MRSA strains started emerging one year after the introduction of methicillin in clinical settings in 1960s [4]. The mecA gene, which encodes the low-affinity penicillin-binding protein 2a causes methicillin resistance in Staphylococci [5]. In recent times MRSA has emerged as an important pathogen of public health importance causing significant morbidity. Most of them are multidrug resistant. Rapid detection of MRSA is essential for early treatment of the patients and implementation of infection control policies to prevent its spread and outbreaks. The nuc gene is specific for S.aureus and molecular detection of this gene helps in the rapid identification of S.aureus from clinical specimens.
There are several phenotypic methods for the detection of methicillin resistance in Staphylococci which includes manual methods and automated systems such as Vitek 2C [6]. The disadvantage of the phenotypic methods are that they are time consuming and have a turnaround time of 18-24 hours.
Molecular methods like Polymerase chain reaction (PCR) and real-time PCR have been used for rapid identification of S. aureus, particularly for MRSA [7,8]. Loop-mediated isothermal amplification (LAMP) assay is a novel method for amplifying DNA and RNA with high specificity, sensitivity, and simplicity. This newly developed tool has enlightened the world of today’s diagnosis and changing the way infectious diseases are monitored in developing countries. Loop mediated isothermal amplification, as the name implies, the reaction takes place at an isothermal temperature, which is the greatest credit to the test. Essentially, the assay consists of incubating a mixture of the target gene along with six sets of target specific primers (2 inner, 2 outer, 2 loop), Bst DNA polymerase and the substrates at an optimal temperature (63—65oC). The inner primers introduce self-complementarity into the amplification product, causing loops to form, while extension of the outer primers causes displacement of the extension products of the internal primers [9].
Compared to PCR, LAMP has the advantages of reaction simplicity and rapidity.
This study was conceptualized with an aim to develop and evaluate the LAMP assay for detection of Staphylococcal spp from cultures using nuc (for identification of S. aureus) and mecA (for detection of methicillin resistance) gene specific LAMP assays.
Materials and Methods
This study was done at Nizam’s Institute of Medical Sciences, Hyderabad from June 2013 to May 2014. The standard control strains used in this study were American Type Culture Collection (ATCC) of 25923 methicillin Sensitive Staphylococcus aureus (positive control for Methicillin Sensitive SA), ATCC 43300 MRSA (positive control for MRSA), ATCC 25922 Escherichia coli (negative control for mecA and nuc genes) (commercially procured from microbiologics, USA). The LAMP assays for mecA and nuc genes were optimized using these ATCC strains, which were later applied to screen the clinical isolates of Staphylococci.
Optimization of the LAMP Assays for mecA and the nuc genes:
Genomic DNA Extraction
Genomic DNA was extracted from overnight growth cultures of all the ATCC strains on 5% sheep blood agar (bioMérieux, Marcy l’Etoile, France) by boiling method, as per the Centers for Disease Control and Prevention (CDC) protocol [10].
Sequence of DNA Target Selection and Primer Designing
mecA and nuc specific gene sequence of different Staphylococcus strains were taken from NCBI site (ncbi.nlm.nih.gov.in) and the sequences were aligned by using clustal W software. After alignment LAMP primers were designed by using highly conserved regions of mecA and nuc genes of Staphylococcal strains with accession no X52593.1 and BA000018.3 respectively, by using Primer Explorer V4 software (http://primerexplorer.jp:81/lam/).
The LAMP primers included two outer primers F3 and B3, two inner primers FIP and BIP, two loop primers (forward loop primer (FLP) and backward loop primer (BLP) (Notomi et al.,). The list of all the LAMP primer sequences selected and used in this study is shown in [Table/Fig-1].
Primer sequences for nuc and mecA genes for PCR and LAMP assay
| Genes | Primer sequences |
|---|
| PCR |
| nucFP | CAAAGCATCAAAAAGGTGTAGAGA |
| nucRP | TTCAATTTTCTTTGCATTTTCTACCA |
| mecAFP | GGCAATATTACCGCACCTCA |
| mecARP | GTCTGCCACTTTCTCCTTGT |
| LAMP assay |
| nuc-F3 | TCGCTTGCTATGATTGTGG |
| nuc-B3 | ACATACGCCAATGTTCTACC |
| nuc-FIP | GTACAGTTTCATGATTCGTCCCGCCATCATTATTGTAGGTGT |
| nuc- BIP | TGTTCAAAGAGTTGTGGATGGTGTACAGGCGTATTCGGTT |
| nuc-FLP | TTGAAAGGACCCGTATGATTCA |
| nuc-BLP | GATACGCCAGAAACGGTGA |
| mecA-F3 | GGTACAAGATGATACCTTCGTT |
| mecA-B3 | ATAGCAGTACCTGAGCCAT |
| mecA-FIP | TCTTCAGAGTTAATGGGACCAAACAGAAAGTCGTAACTATCCTC |
| mecA-BIP | AAGCTCCAACATGAAGATGGCTTGTATGTGCGATTGTATTGC |
| mecA-FLP | ACCTAATAGATGTGAAGTCGCT |
| mecA-BLP | CGTGTCACAATCGTTGACG |
FP-Forward primer
RP-Reverse primer
F3-Forward outer primer, B3-Backward outer primer
FIP-Forward inner primer, BIP-Backward inner primer
FLP-forward loop primer, BLP-backward loop primers
All the LAMP primers were synthesized commercially (Bioserve, Hyderabad).
Conventional PCR Assay
A 210 bp region of mecA and 90 bpregion of nuc genes were tested by conventional PCR assay. Published primers [11] used in the study were shown in [Table/Fig-1] (Bioserve Technologies, Hyderabad). The bacterial genomic DNA extracted was used as template for amplification. PCR was done on GeneAmp PCR system 2400 (Applied Biosystems, USA) with quick load master mix (New England Biolabs, UK). PCR conditions were maintained at 950C for 10 minutes, initial denaturation followed by 40 cycles of 95 0C 15 seconds, 580C for 30 seconds and 700C for 30 seconds and final extension 700C for 2 min thermal cycler (Perkin Elmer, USA). The PCR products were tested for nuc and mecA genes by running agarose gel electrophoresis. The bands were visualized by the Gel documentation system, (Syngene, UK).
LAMP Reactions
LAMP reactions were carried out as per manufacturer’s instructions (DNA amplification kit; Eiken Chemical Co. Japan). The reaction was standardized at 60oC for 60 min and inactivated at 80oC for 5 min in a water bath.
Detection of LAMP products was by: 1) visual fluorescence by adding 0.2μl of 1/10000 DMSO SYBR Green I (Sigma Aldrich, USA) to 25 μl of LAMP product; 2) A 10μl of the LAMP product was electrophoresed on 2% agarose gel and documented using a gel documentation system (Syngene, UK).
Clinical Isolates
A total of 100 (70 from purulent aspirates of deep seated abscesses and postoperative wounds and 30 from blood cultures), clinically significant, non-duplicated Staphylococcal isolates were included in this study. Phenotypic identification and antimicrobial susceptibility of the isolates were carried out with the Vitek 2 system, using the ID GP and the P628 AST panels (bioMerieux, Marcy l’Etoile, France). Vitek 2 Advanced Expert System (AES) analysis findings were recorded.
The optimized LAMP assays and conventional PCR for the 2 genes were applied to screen these 100 clinical isolates of Staphylococci.
The phenotypic, conventional PCR and LAMP assays were compared for the identification and detection of clinical isolates of MRSA. The turnaround times (TAT) and cost per test were calculated for these methods.
Results
The specificity of the LAMP F3 and B3 primers were confirmed by PCR on the ATCC strains (positive and negative controls). [Table/Fig-2] shows the agarose gel image of the PCR result of the nuc and mecA genes, found only in the ATCC standard for MRSA and the nuc gene alone in the ATCC MSSA.
nuc and mecA conventional PCR agarose gel image by using LAMP outer (F3 B3) primers

The specificity of the LAMP assays for the 2 genes were 100% on the ATCC strains.
Among the 100 Staphylococcus isolates, the Vitek 2 identified 82(82%) isolates as S.aureus and 18(18%) isolates as CONS and detected methicillin resistance in 56 isolates (44 MRSA and 12 MRCONS)
The conventional PCR and LAMP assays also showed identical results. However, the results of the LAMP assay were rapid and were available within 90 minutes as compared to the Vitek 2 results (18-24 hours)
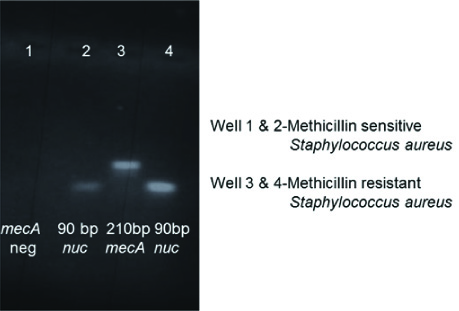
The gel images of conventional PCR using published forward and reverse primers are shown in [Table/Fig-3]. The visual fluorescence of the LAMP products and the gel images are shown in [Table/Fig-4,5]. The amplified target sequences of LAMP assays can be visualized as ladder like bands in the gel image due to the formation of stem loop structures of amplified DNAs of various stem lengths.
nuc and mecA conventional PCR agarose gel image by using published primers
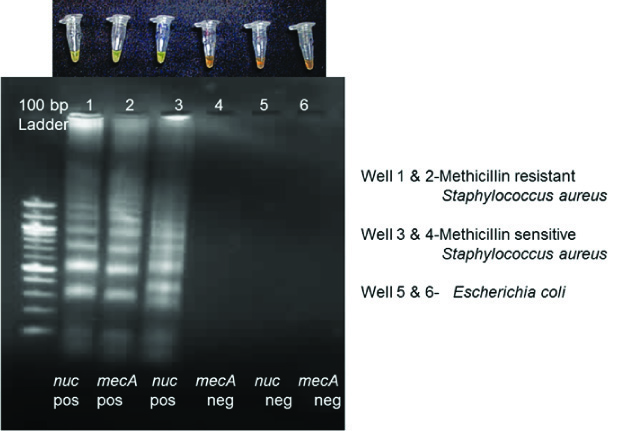
Visual fluorescence and gel images of LAMP products of MRSA and MSSA
Visual fluorescence and gel images of LAMP products of MRCONS and MSCONS

The comparison of TATs and cost per test of phenotypic, conventional PCR and LAMP assays are shown in [Table/Fig-6].
Comparison of Turnaround time and cost per test for phenotypic, conventional PCR and LAMP Assay

Discussion
S. aureus, including MRSA, is one of the most important bacteria causing a wide spectrum of human infections [12]. Timely detection and confirmation of MRSA is important for adequate and appropriate treatment, surveillance and infection control.
There are very few studies using LAMP assays for the detection of Staphylococcal isolates, which include femA and mecA genes from food isolates [13], femA from clinical and food samples [14], spa and arc C genes from spiked blood specimens [7], spa and mecA gene from positive blood culture bottles [15], orfX (a highly conserved open reading frame of S.aureus) from clinical isolates [16].
In this study, we differentiated S.aureus from CONS using the nucgene. S. aureus strains produce an extracellular thermostable nuclease (thermonuclease {TNase}), which is an endonuclease that degrades both DNA and RNA characterized by its gene nuc specific for S.aureus [17,18].
Methicillin resistance was detected by mecA gene in both S.aureus and CONS. mecA gene, specific for Methicillin resistance, is carried by a mobile genetic element Staphylococcal cassette chromosome mec (SCCmec) is characterized by the presence mecA gene in mec gene complex,ccrgenes(ccrA and ccrB) in ccrgene complex, and the presence of flanking direct repeat sequences containing the integration site sequence (ISS) [19,20].
In comparison to phenotypic methods, which are time consuming and PCR, which requires thermal cycler and requires technical expertise, LAMP uses a simple water bath and allows visual detection of amplified product by turbidity or fluorescence. The LAMP assay is a more rapid and simpler method than PCR. The total amplification time for LAMP is 60 minutes, as compared to PCR, which is about 3-4 hours. It is based on autocycling strand displacement DNA synthesis using the Bst DNA polymerase enzyme. It uses six primers recognizing 6—8 distinct regions of the target gene facilitating the detection of very minute quantities of target DNA. The amplification occurs under isothermal conditions between 63°C [9,14] and time loss due to thermal changes is prevented [7]. In addition the reaction is not inhibited by the inhibitors in the sample [21]. The amplified gene product can be visualized by the unaided eye, either as turbidity in the form of a white precipitate or through a colour change employing a fluorescent intercalating dye (SYBR Green I) [9]. The results of LAMP assay was comparable with phenotypic and conventional PCR with the advantage that the LAMP assay is rapid with a turnaround time of 90 minutes following the isolation of the organism from clinical specimen and is also cost effective.
Conclusion
The present study proved that LAMP assay can be used as a rapid molecular diagnostic assay for the simultaneous differentiation of Staphylococcal spp and detection of methicillin resistance. It can be easily adapted in any microbiology laboratory as a rapid molecular bench side assay due to its simplicity and ease of performance without any sophisticated instrumentation.